 In comparison, the other file formats are geared towards data visualization and dissemination, thus their specifications aim to facilitate user-friendliness.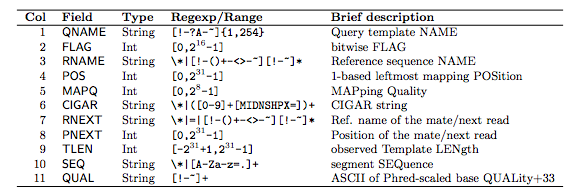 
The consortium considers FASTQ as the basic file format for archival purpose and thus the FASTQ format's specifications aim to preserve the raw sequence data. As the ENCODE consortium is a collaborative effort, the consortium has made several specifications on the file formats to facilitate data archival, presentation, and distribution, as well as integrative analysis on the data. These file formats were originally designed to be generic and flexible. hic: The hic format is a binary format for storing contact matrices and annotations of chromatin structural features generated from Hi-C or other proximity mapping assays.bigBed: The bigBed format is also an indexed binary format for rapid display of annotation items such as a linked collection of exons or the binding peaks of a transcription factor.bigWig: The bigWig format is an indexed binary format for rapid display of continuous and dense data in the UCSC Genome Browser.BAM: The Sequence Alignment/Mapping (SAM) format is a text-based format for storing read alignments against reference sequences and it is interconvertible with the binary BAM format.FASTQ: a text-based format for storing nucleotide sequences (reads) and their quality scores.The ENCODE consortium uses several file formats to store, display, and disseminate data: XN: Amplicon name tag, which records the amplicon tile ID associated with the read.īAM index files (*.bam.bai) provide an index of the corresponding BAM file.Common File Formats Used by the ENCODE Consortium Overview NM: Edit distance tag, which records the Levenshtein distance between the read and the reference. RG: Read group, which indicates the number of reads for a specific sample.īC: Barcode tag, which indicates the demultiplexed sample ID associated with the read. The alignments section includes the following information for each read or read pair: The read name includes the chromosome, start coordinate, alignment quality, and the match descriptor string. 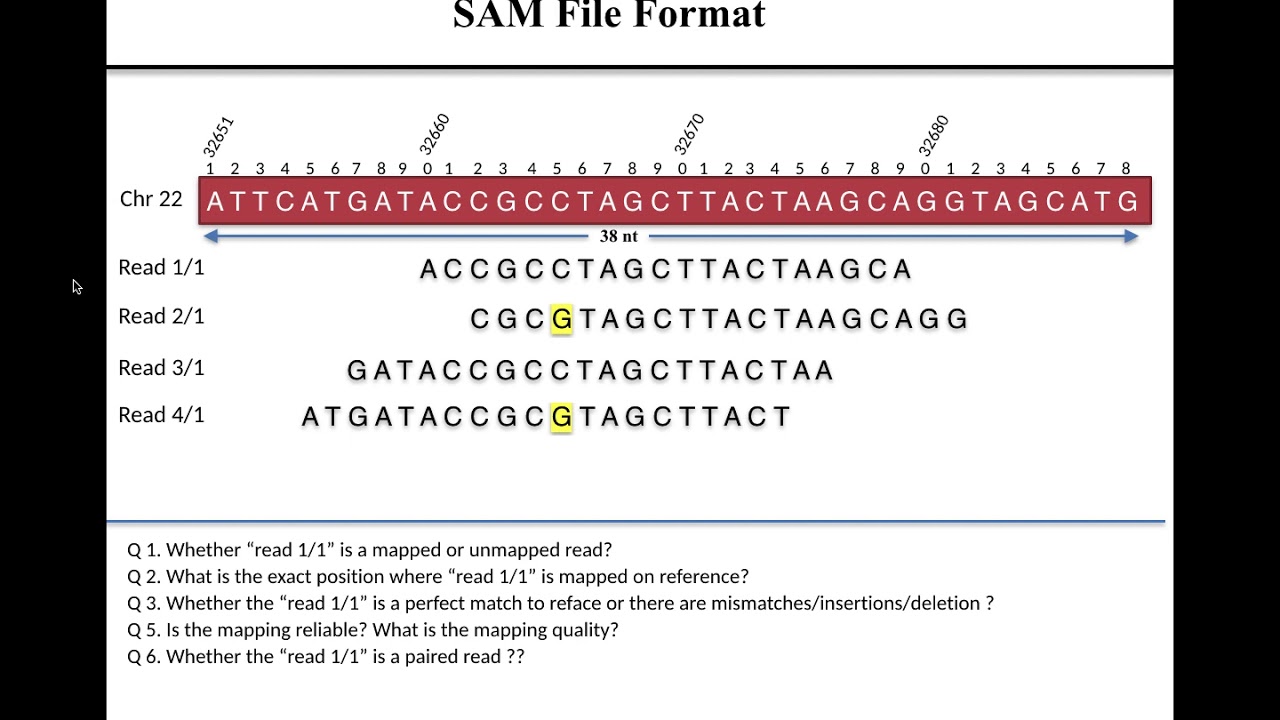
Alignments in the alignments section are associated with specific information in the header section.Īlignments-Contains read name, read sequence, read quality, alignment information, and custom tags. Header-Contains information about the entire file, such as sample name, sample length, and alignment method. Whole Genome Sequencing v5.0 is multinode only and uses the file naming format SampleName_S1.bam.īAM files contain a header section and an alignment section: 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
